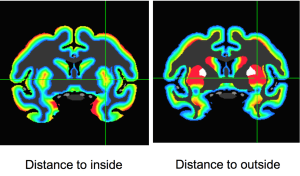
There is a long list of software packages that are capable of performing high-resolution MRI analysis.
Some of them are used by multiple groups and some of them are customized for specific studies only.
In this post, I want to give an overview over the most important software packages, their advantages and disadvantages, and their popularity in the field.
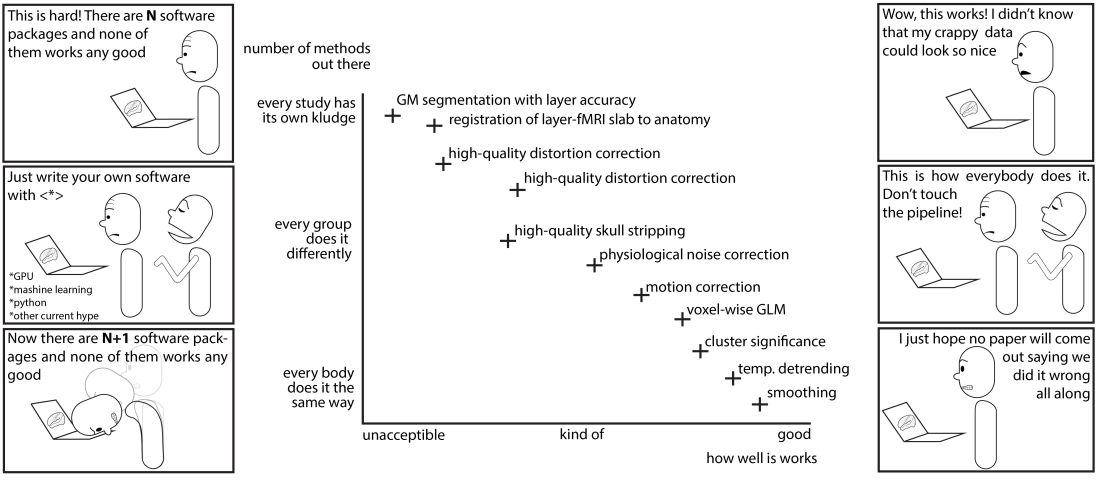
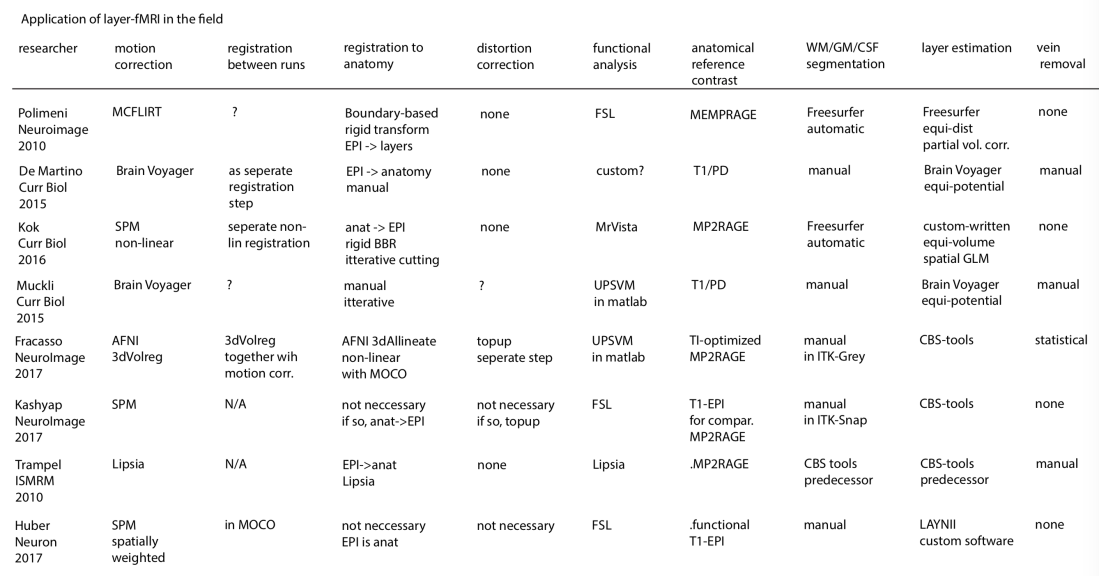
Analysis tools applied in the seminal layer-fMRI studies
Freesurfer
Description: Freesurfer is an open source software suite for processing and analyzing (human) brain MRI images. In the recent years, it was further optimized for ultra-high-resolution data. In the field of layer-fMRI it is the most often applied software package.
Advantage: The usage is very well documented with an extensive Wiki. It is very robust and has a very large user-community.
Disadvantage: It takes quite some time. Some people say it would have a rigid data organization. As a user, it’s not so straight forward to adapt/understand the algorithms and/or build your own tools. It requires a whole brain coverage, while there is not whole brain layer-fMRI acquisition protocol out there (yet). It is not straightforward to manually correct segmentation errors with a sub-voxel accuracy.
Dependencies: Stand alone, no dependencies. It works well on Unix and Mac.
Link: https://surfer.nmr.mgh.harvard.edu/
A layer-fMRI pipeline using Freesurfer in matlab was developed by Kendrick Kay.
My issues with it
I ended up not using it because I found it very hard to include manual corrections in the segmentation. It never works perfectly (especially in M1) and manual corrections can only be made on an integer voxel level. Dependent on the brain area segmentation does not always work perfectly (like in all software packages) and manual corrections are inevitable.
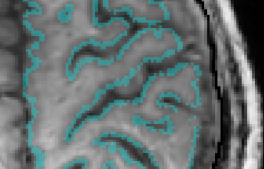
Freesurfer does the layering based on the WM/GM surface. This results in larger errors and non-linearities towards the cortical surface.

Without further ado, it provides surfaces as meshes in standard space. I never got it to work that I obtain surfaces in voxel space with the original oblique NIFTI orientation.
The mesh density at the surface is very irregular and often less than the voxel size, this means that it misses many voxels.


CBS-tools

Description: CBS High-Res Brain Processing Tools (CBS Tools) is a software suite optimized for processing of MR images at sub-millimeter resolutions. CBS Tools have been developed in Java as a set of plugins for the MIPAV software package and are controllable from the JIST pipeline environment.
Advantage: It is optimized for layer-fMRI and has the equi-volume layering algorithm included.
Disadvantage: Complicated to install and use (for me). Is requires whole brain data for the segmentation. The layering works with partial brain data, however.
Dependencies: It is used as a collection of plug-ins in MIPAV.
Link: https://www.nitrc.org/projects/cbs-tools/
My issues with it
I never got it to work on my machine. This is so despite support from former developers.
Nighres
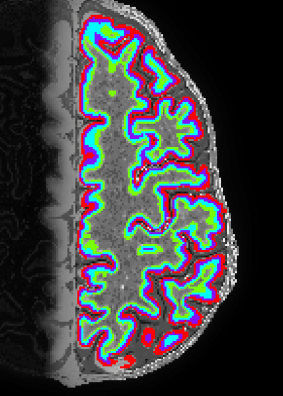
Description: Nighres is a software tool in Python that is built on CBS tools developed by Julia Huntenburg, Christopher Steel and Pierre-Loiuse Bazin. It is specifically designed for high-resolution, layer-dependent MRI analyses.
Advantage: Its open source and very well documented. Its super fast. On my laptop it takes 15 min to get from MP2RAGE data to layers in volume space.
Disadvantage: It is not easily usable on MacOS, just yet. It requires whole brain data for the application of segmentation (it uses whole brain reference atlases). Layering can be done with partial coverage for closed surfaces. Even though it is designed to allow changes made by users, the complicated meta structure of multiple wrappers across multiple programming languages requires advances coding experience.
Dependencies: wrap, JCC, Nimpy, Nibabel
Link: http://nighres.readthedocs.io/en/latest/installation.html
My issues with it
Because I am forced to use a MAC, I can only run it in a Neurodebian virtual box. This makes it more complicated for me. I also found it quite RAM-hungry. I guess I will use it more, when it will be easier available for MAC and when there are more functions being included.
AFNI
The AFNI suite is also often used in layer fMRI for few sub-steps along the evaluation pipeline.
Afni can be used to isolate the voxels of a given cortical depth with 3dSurf2Vol using Freesurfer GM and WM surfaces as input. Note the arguments –f_p1_fr and -f_pn_fr that defines the range of cortical depth. See an example of how this works in this previous blog post.
Daniel Glen presented a poster at HBM 2018 showing multiple new tools in AFNI that estimate cortical thickness. These algorithms can also be used as a measure of cortical depth.
Advantage: As most other AFNI tools, the layer-tools are also fast and mighty.
Disadvantage: The segmentation is usually obtained form Freesurfer anyway, so it appears natural to do the layering estimation in directly in free surfer too. One thing that I found particularly challenging and that keeps me for applying them on a more regular basis is the problem that it needs some extra work around to work with oblique slices (see line with 3dWarp -card2oblique EPI.nii -verb anat.nii orinentfile.txt in this example)
Dependencies: The installation of AFNI required quite a bit of preparation work and other software to be installed. But it is quite well documented and it succeeded doing it few times already.
Link: https://afni.nimh.nih.gov/
LAYNII
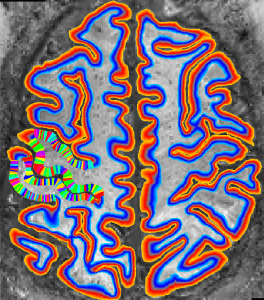
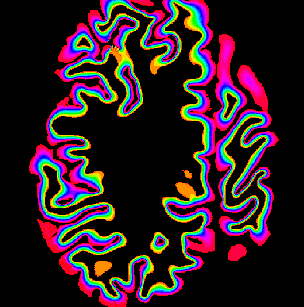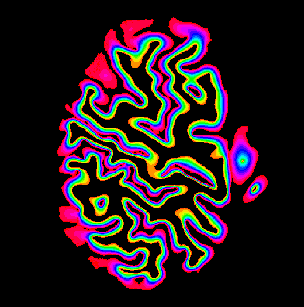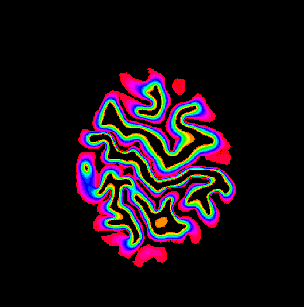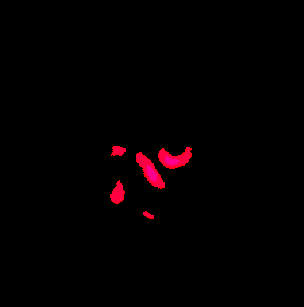
Description: Collection of C++ programs developed from Renzo Huber that operate exlusively in voxel-space.
Advantage: Because I wrote it myself, I know what it does and how the algorithms work 😉 Very flexible and application for real-life data with limited FOV. Since its implemented in C++, its super fast.
Disadvantage: Some functions are using ODIN libraries for the reading and writing of Nifties, which requires a unix machine with root access to be installed This requirement is not easily fulfilled in all labs
Dependencies: Old programs use ODIN libraries for reading and writing nifties. This will hopefully change in the future. Few programs use GSL.
Link: https://github.com/layerfMRI/LAYNII (without ODIN dependencies) and https://github.com/layerfMRI/repository (with ODIN dependecies).
My issues with it
I didn’t transfer all the programs to a NIFTI I/O to a standalone C++, just yet. Hence, many programs still depend on ODIN libraries and are only applicable on UNIX machines. The equi-volume layering is still not optimized for oblique slices that are curved in 3D.
It does not have a very robust tissue segmentation algorithm, yet. So I need to use alternative methods and manual corrections.
Brain Voyager
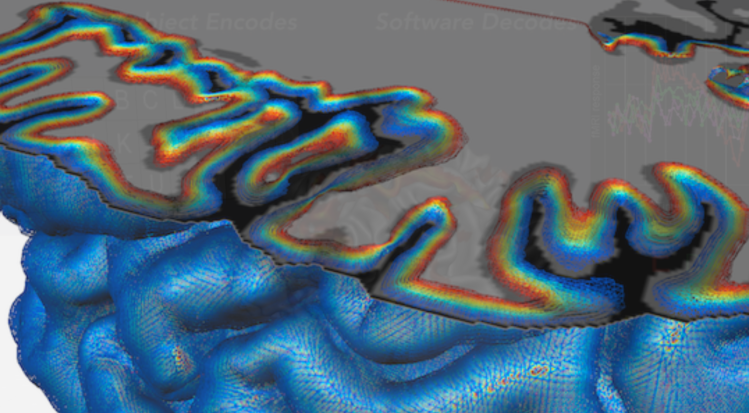

Description: Non-open source software developed around Reiner Goebel for MRI analysis. It has a specific focus on laminar and columnar analysis.
Advantage: The 3D-grids to pool voxels across layers and columns is particularly advantageous. A nice description of all the layer-fMRI features are nicely explained by Valentin Kemper in this recent paper:
Disadvantage: Expensive. Its a black box.
Link: http://www.brainvoyager.com/
My issues with it
Since it is not open source and requires non-cheap licenses, I never used it myself.
Segmentator
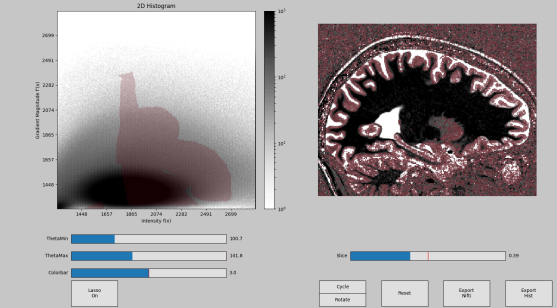

Description: Segmentor is an open and freely available Python tool box written by Faruk Gulban and co-developed by Marian Schneider for tissue type classification in high-resolution (layer)-fMRI. Segmentator plots voxels in a 2D-histogram. Every voxels’ signal intensity is plotes vs. its local gradient. Segmentator algorithm take advantage of the fact that non-brain structures (vessels, dura, CSF) have a spatial extent of approximately 0.7mm, which is coincidentally the current limit of routine anatomical images at high fields. Hence, non-brain voxels can be nicely separated from brain voxels in the gradient space.
Advantages:
- it’s faster than manual segmentation and a first step for manual correction
- is can work with any volume data-set larger than 3x3x3 voxels.
- it works with slab-data, surface-coil data, and it doesn’t require whole brain
- it can be optimized for any desired small brain region of interest.
- its relatively easy to install.
- its a tool that can be inserted in any other analysis pipeline.
- it provides tools that are not included in any alternative repository.
- it takes advantage of multiple images/contrasts at once.
Disadvantage:
- it neglects valuable information on a more larger spatial scale: e.g. atlas based segmentation priors, additional conditions about GM ribbon smoothness, etc.
- it is only a first step toward fully-automated segmentation and still needs to be applied in combination with other methods
Dependencies: Python, pip for installation, Matplotlib, Numpy, Nibabel, Scipy
Link: https://github.com/ofgulban/segmentator
My issues with it
It is only a supporting tool that needs to be applied in combination with other methods. E.g. manual refinements are still often necessary. It requires relatively high quality data.
RBR (Recursive Boundary Registration)
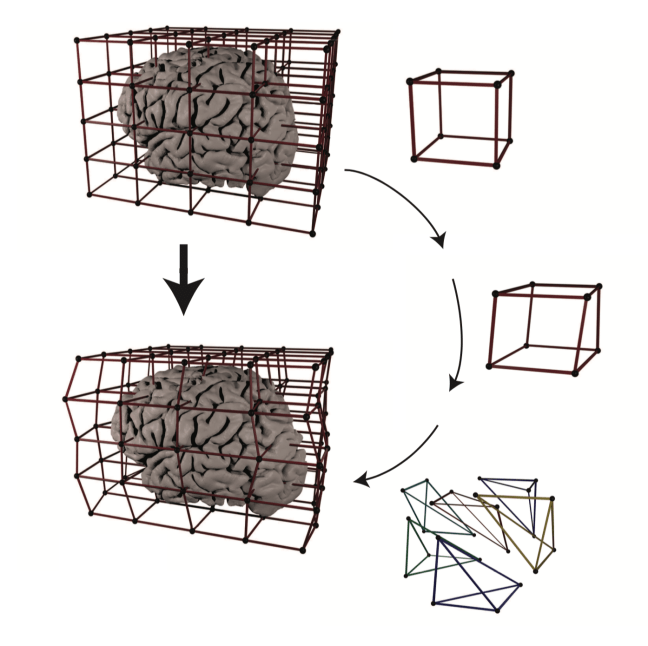
RBR is a Matlab suite developed from Tim van Mourik for high-quality non-linear registration between undistorted ‘anatomical’ data and functional (layer) fMRI data. Its algorithm is based on repeatedly cutting the brain in smaller and smaller chunks and registering them with higher quality as it would be possible with larger volumes.
Its is comprehensively described in a resent bioarchive paper https://www.biorxiv.org/content/early/2018/01/15/248120.
Advantage: It has a higher registration quality than whole-volume registration.
Disadvantage: There is a tradeoff between conserved topology and smoothness of displacement that can restrict the registration quality.
Matlab code: https://github.com/TimVanMourik/OpenFmriAnalysis
My issues with it
It is not clear to me how it performs with respect to other methods that are optimized for large distortions. E.g. using a less conservative smoothness penalty in the deformation field of SyN (in ANTS: SyN[0.1,-> x <-,0]). Field distortions are also not so variable across time that TopUp is completely useless. Any within-run registration would be impossible too.
Herrlich10 tools
Github repository to extract equi-volume layers from vertices with Python: Paper: https://doi.org/10.1101/2020.02.01.926303
code: https://github.com/herrlich10/mripy/blob/master/mripy/scripts/mripy_compute_depth.ipy
Pipeline of CVN-Lab
This is the pipeline for developed from Kendrick Kay and his lab members for layer-fMRI. It consists of a whole set of layer-fMRI preprocessing tools including: alignment, field-map calculation, distortion-correction, registration, functional analysis and layering. It is written in matlab and called multiple external programs. As such, it uses freesurfer for the segmentation and layering.
Advantage: The quality seems to be exceptional. Kendrick’s paper seems to be one of very few that achieve usable segmentation quality without too much manual intervention. The visualisation tools are very instructive and give the user a good feeling on how the data are affected as the go through the evaluation pipeline.
Disadvantage: It is not written in a way to be use from a large user group. I have the feeling that you need to be Kendrick himself to set it up and run is robustly. The repository can be useful to copy code snippets, though.
Matlab code: https://github.com/kendrickkay/cvncode
Surface tools
Description: Collection of tools for surface-based operations developed from Konrad Wagstyl, Casey Paquola, Richard Bethlehem, Alan C Evans, and Alexander Huth written in python.
It can use Freesurfer surfaces and to estimate equi-distant and equi-volume surfaces across the cortical depth.
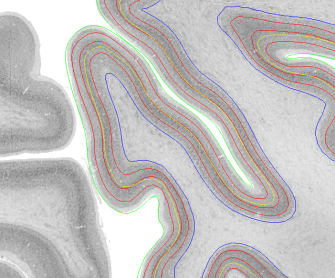

Code and instructions: https://github.com/kwagstyl/surface_tools
Biggest unresolved issues in layer-fMRI analysis
Registration quality on a sub-voxel accuracy basis
There is no established quantitative measure that provides an estimate of registration quality up to a accuracy level of cortical layers. There are many suggestions, but its not clear which one performs best and whether any methods out there works good enough at all.
- Tim van Mourik suggests iterative BBR on smaller and smaller junks of the brain
- SyNs in ANTs (me)
- 3dQwarp (Yuhui)
- SPM (Yinghua)
I think future work should test which of the methods works best. Future work could be focusing on a good parameter set with the best accuracy-efficiency tradeoff. Furthermore future stream-lining work could try to use warp feels across platforms so it can be combined with motion correction.
Segmentation with sub-voxel accuracy
There are multiple ways of doing segmentation and it is not clear which one is the best.
- I start with Freesurfer (highres option) segmentation and do a manual corrections.
- Running 3dkMeans in AFNI like Allessio Francasso. It works better when you run 3dKmeans for different chunks of the brain separately and then merge them afterwards.
- Segmentator from Faruk
- ITK-snap with its growing balloon algorythm
- Using texture and local entropy like John Schwarz
- SPM and manual adjusted “winner-map”
I think future work should go into the direction of finding out which of those methods has the best accuracy-efficieny tradeoff.
Alignment without resolution loss
Every resampling step results in spatial blurring. It also introduces spurious correlations of voxels the time series. There have been three suggestions to account for it:
1.) Apply all registrations in one single resampling step only. Motion correction, alignment of runs, distortion correction, alignment with anatomical reference. As far as I understand. Nobody it using this approach so far. It is not straight forward to combine them in one step because different software packages are used for all those steps.
2.) Only do motion correction in EPI space. And ally the remaining analysis is done in EPI-space. This is what I do. However, freesurfer and AFNI are not making it easy to work in subject-specific distorted space. Future developments should make this easier.
3.) Up-sample all coil-data in k-space at the scanner and apply the analysis on a finder grid than the effective resolution. This approach means that every dataset is now 2^3=8 times larger. Which makes the analysis longer, memory requirements tougher. Data-analysis cannot be done on local laptops anymore and. Data transfer to HPC resources takes hours and is impractical.
I think it is still not established how bad multiple resampling steps reduce the resolution. I think future work should compare different pipelines and see if the layer-peaks are particularly clear for one or the other.
Segmentation of T1-EPI data
T1-EPI has more artifacts than MP2RAGE data. And it usually does not have whole brain coverage, which makes it harder to use it in conventional segmentation and layering pipelines. Since it also has a lower CNR, it requires more manual interventions. It is not established how to preprocess T1-EPI data so conventional segmentation pipelines can deal with them more efficiently.
Streamlining of manual corrections
So far, every respectable layer-fMRI paper does some sort of manual correction. This often happens during the alignment step and/or during GM segmentation.
There are multiple ways how to do manual correction:
- ITK-snap allows users to do manual segmentation with inflating balloons. In my opinion it’s quite time consuming.
- FSL allows me in my unscaled layer space to shift border lines by 25% of a voxel size. Is takes about 3 hours to do manual corrections as long as I have a good starting point from freesurfer.
- Freesurfer has its own way of doing manual correction on an integer voxel basis. Hence, layer position accuracy cannot be better than the voxel size.
I think that future work should go into the direction of combining independent approaches (Segmentator, Freesurfer, 3dclust) to obtain a better starting point for manual corrections.
I think future pipelines should be streamlined to allow manual corrections and it should become easier to incorporate manual corrections into the pipelines.