Almost every modern fMRI protocol (at SIEMENS scanners) uses GRAPPA. However, only very few people pay a lot of attention on optimal usage of the GRAPPA auto-callibration data. I realized the importance of optimizing GRAPPA parameters when doing high-resolution EPI. At high resolutions, GRAPPA-related noise can become an increasingly important limitation. This is especially true with the low bandwidth that the body gradient coils force us to use.
In this blog-post I will explain how the GRAPPA kernel-size affects the fMRI data quality, how you can change it, how you can find out which kernel-size was used, and I will descrive simple software tools to identify regions that might benefit from adaptations of the GRAPPA-kernel size.
Effect of GRAPPA kernel-size
The GRAPPA kernel is used to synthesize missing k-space lines in accelerated acquisitions. Its exact working principles are a bit technical. If you are interested in the full mathematical model, I found the original GRAPPA paper quite comprehensive and understandable.
In my simplified understanding of the working principle of GRAPPA, the GRAPPA kernel defines the size of the spacial smoothness of the coil-sensitivity maps. This comes from the k-space feature that small objects in k-space (small kernels) are associated with smooth objects in image space. Larger objects in k-space (larger kernels), on the other hand, are associated with sharper objects in image space.
Keeping this in mind, the GRAPPA-kernel size must have some effect on the spacial distribution of the g-factor penalty.
Namely, smaller GRAPPA kernels will have smoother defined regions of the high g-factor penalty
Small kernel example:
Large kernel example :
The corresponding temporal stability is apparent in tSNR maps.



The GRAPPA parameter of the data shown are: 7T, resol. = 0.75mm, phase-encoding direction = A>>P, GRAPPA-factor = 3, ACS-scheme = FLASH (30 dummies, FA 5deg, BWDTH = 100 Hz/px), #ref lines = 48, FOV = 130 mm, Coil = 32ch Nova.
As you can see, in my data, the GRAPPA-kernel size has an effect of about 20-30% improvement/reduction in the final tSNR.
When the area of interest is suffering from large g-factor penalties (usually close to the center of the FOV), I would recommend a small GRAPPA kernel. When the area of interest is just outside the area of large g-factor penalties, I would use larger kernels.
This, of course is highly dependent on the FOV, the GRAPPA factor, the CAIPI FOV-shift, and the size of the RF coils. As such, for smaller RF-coil elements (e.g. in surface arrays, high-channel count arrays) or very large FOV, larger GRAPPA-kernal sized would be recommended.
It is also worth pointing out that the GRAPPA-kernel-size also affects the reconstruction speed. Larger kernels result in longer reconstruction durations. In my, experiments however, I am not really limited by reconstruction speed. So, I done mind. practiCal compared the reconstruction durations more rigorously in his blog.
How to change the kernel-size on a VB17 SIEMENS system
There are multiple ways how you can change the GRAPPA-kernel size on a SIEMENS VB17 scanner. Below, I describe the most common approaches.
Adjustment by changing Ice-Configurations (recommended)
- Start “xbuilder” (Ctrl+Esc and run xbuilder)
- Open the file c:/MedCom/config/Ice/IceConfig.evp
- Find parameter ICE/CONFIG/iPAT/ImprovedGrappa and set it to “1”

- Find ICE/CONFIG/iPAT/GrappaPSize or ICE/CONFIG/iPAT/GrappaSegements just below and change them as you wish. E.g. change GrappaPSize from 4 to 2.

Note: The file IceConfig.evp is overwritten every time a new ‘patient’ is registered. Hence, if you mess around in IceConfig.evp, there is no danger that you affect other people scanning after you. However, it also means that you should always apply your changes after your patient is registered.
Note: The GRAPPA-kernel size in the current IceConfig.evp file is also used for Retro-Recon. This means that when you want to RetroRecon an old dataset while a different patient is currently registered. The setup of the current patient will be used. This feature also allows you to reconstruct the same date with multiple GRAPPA-kernel sizes. Just change IceConfig.evp in the current patient and any data will be reconstructed with the current set of parameters during RetroRecon in Twix.
Adjustment by setting it in the Ice-Program
The values within the IceConfig.evp can be overruled in the IceProgram. I.e. if you use your own IceProgram, you can include additional changes as follows:

I do not really have good experiences with this way of doing it. I found that it sometimes fails (in RetroRecon only) when there are conflicting parameters in IceConfig.evp and the IceProgram*.evp. Markus Barth told me that this Retro-Recon problem can be avoided when the corresponding “EpiGrappaParameters” field is deleted from the twix file (the the one at the bottom).
Adjustment within the sequence special card
Some WIP sequences allow you to adjust the kernel-size via the sequence special card. E.g. the CMRR MutliBand sequence allows you to choose 3×3, 5×3 and 5×5 (slice-size x read-size).
How to find out which kernel-size was eventually used
Very often you would like to find out which GRAPPA-Kernel size was used. And/or you want to double check whether your manually changed kernel-size was properly applied. You can check the used GRAPPA-kernel size in the Twix data as follows:
- Start Twix (Ctrl+Esc and run twix)

- Select the dataset in the left panel (red), open RetroRecon (blue), and click on edit (green).
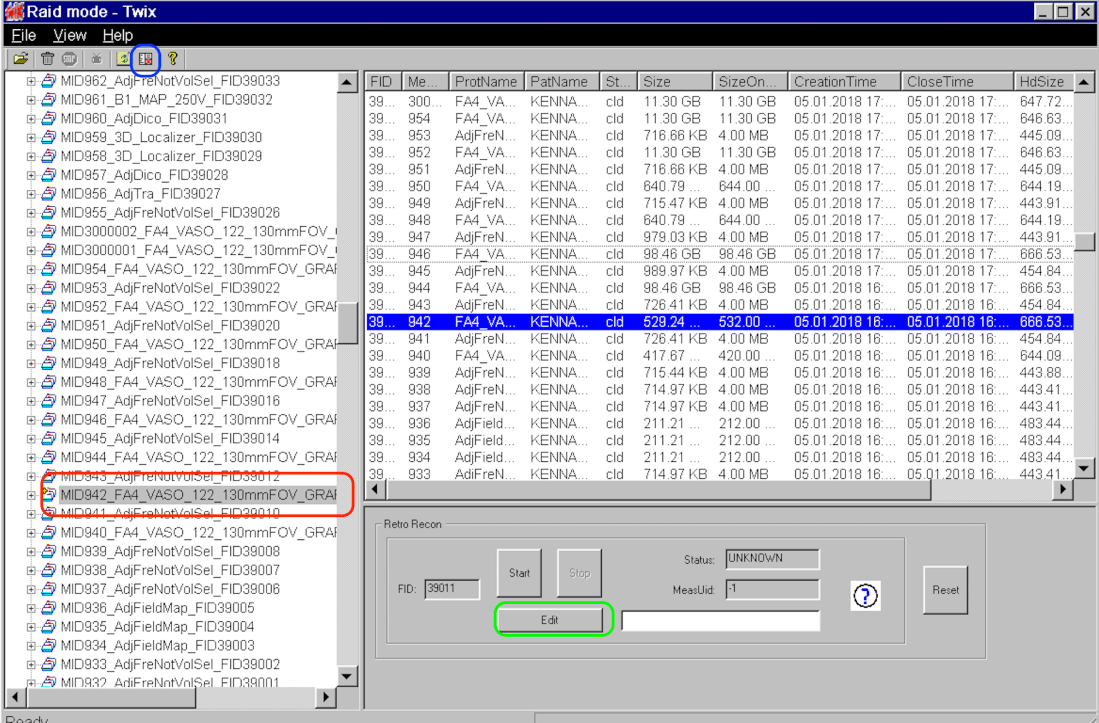
- The used GRAPPA-kernel size can be seen in -> XProtocol-> IRIS -> PIPI -> GrappaFindWs -> Functor (black arrow).

Simple software to explore optimal GRAPPA-kernel sizes for a given ROI
The program LN_GFACTOR is part of the standalone C++ software suite LAYNII. Its whole purpose is to find regions of particularly high g-factor penalty for a given dataset, GRAPPA-factor, phase-encoding direction.
It can be called with the following command:
LN_GFACTOR -input input.nii -output GFACTOR.nii -variance 1 -direction 1 -grappa 2 -cutoff 150
It will provide outputs as shown above. G-factor maps and simulated noise distribution.
Terminology of GRAPPA Kernel size
In SIEMENS jargon, the Kernel size is often provided in the terminology as follows: 3×2 (read-size x phase-size). The in “IceConfig.evp” they are referred to as “GrappaPSize” = 2 and “GrappaSegments” = 3.

Since the reconstructed data are not acquired in in-plane GRAPP the Phase encoding size must be an even number. Since all data are acquired in read direction the GRAPPA kernel in read direction must be an odd number.
Many people are coming in contact with Kernel sizes with the CMRR MultiBand sequence. This sequence allows the user to choose the Slice-GRAPPA Kernel size in the special card. Note however that the slice-GRAPPA Kernel is a different Kernel than the In-plane GRAPPA-Kernel. E.g. you can have a in-plane GRAPPA kernel of 3×2, and at the same time you can have a slice-GRAPPA kernel of 5×5 (read-direction x slice direction). E.g. see the examples in the supplementary material of a recent paper that can be downloaded here (Figs. S2-S5). There we found that with conventional 32ch coils, smaller kernels result in better fMRI stability.
In slice-GRAPPA reconstructions, there are no missing data that need to be filled in. Instead, all the slices are acquired and just need to be unaliased. Hence, for slice-GRAPPA both dimensions of the GRAPPA-Kernel need to be an odd number.



