Overview
Big steps:
- Manual aligment of MP2RAGE with EPI (optional when MP2RAGE is acquired in same session)
- ANTS alignment of MP2RAGE and EPI. (part of anatomical_maser.sh, see github)
- Running Freesurfer on MP2RAGE data in EPI space. (part of anatomical_maser.sh, see github)
- Using SUMA to get fine samples tissue borders in EPI-voxel space (in oblique space) (part of anatomical_maser.sh, see github) Thanks to Daniel Glen developing this workflow.
- Manual correction of Freesurfer GM-ribbon
- calculating layers from GM-ribbon in neuroDebian.
Required:
- whole brain MP2RAGE/MPRAGE
- slab T1-weigted EPI map
- Software: ITK Snap, ANTs, Freesurfer, AFNI, SUMA, Neurodebian VirtualBox from Renzo: https://nimhactivecho.nimh.nih.gov/t/2whrezbd
Output:
- fine scale EPI data up sampled
- Equi-distance Layers in EPI space
1.) first manual alignment (not always necessary)
you need ITK-SNAP (news as 3.6.0): http://www.itksnap.org/pmwiki/pmwiki.php
- open T1-EPI
- Add another image MP2RAGE (semi-transparent) “w” shifts back and forth
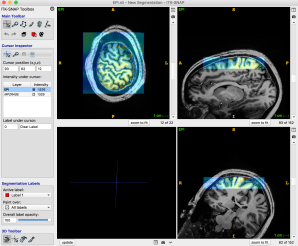
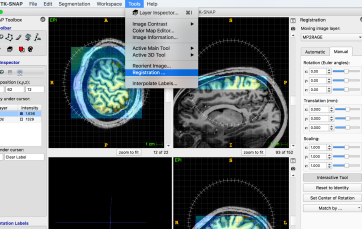
- Select: Tools -> Registration -> Manual
- adjust approximately manual
- you go to automatic tab and run Registration. I liked the following parameters:
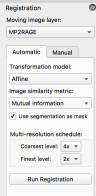 However, note that this does not always work. When the FOV is too small, the automatic registration puts the brain out of place. In this case, the manual adjustments need to be suffice.
However, note that this does not always work. When the FOV is too small, the automatic registration puts the brain out of place. In this case, the manual adjustments need to be suffice. - save the transformation file: with safe symbol

- Now you can apply it with command:
-
antsApplyTransforms --interpolation BSpline[5] -d 3 -i MP2RAGE.nii -r EPI.nii -t initial_matrix.txt -o registered_applied.nii
ANTS documentation: https://github.com/stnava/ANTsDoc/raw/master/ants2.pdf
2.) Application of initial alignment in ANTS
echo "I expect 2 filed. the T1_weighted EPI.nii and a MP2RAGE_orig.nii" # bet MP2RAGE_orig.nii MP2RAGE.nii -f 0.05 3dcalc -a MP2RAGE.nii -datum short -expr 'a' -prefix MP2RAGE.nii -overwrite ITK_GLOBAL_DEFAULT_NUMBER_OF_THREADS=50 export ITK_GLOBAL_DEFAULT_NUMBER_OF_THREADS echo "*****************************************" echo "************* starting with ANTS ********" echo "*****************************************" #2 steps antsRegistration \ --verbose 1 \ --dimensionality 3 \ --float 1 \ --output [registered_,registered_Warped.nii.gz,registered_InverseWarped.nii.gz] \ --interpolation Linear \ --use-histogram-matching 0 \ --winsorize-image-intensities [0.005,0.995] \ --initial-moving-transform initial_matrix.txt \ --transform Rigid[0.05] \ --metric CC[EPI.nii,MP2RAGE.nii,0.7,32,Regular,0.1] \ --convergence [1000x500,1e-6,10] \ --shrink-factors 2x1 \ --smoothing-sigmas 1x0vox \ --transform Affine[0.1] \ --metric MI[EPI.nii,MP2RAGE.nii,0.7,32,Regular,0.1] \ --convergence [1000x500,1e-6,10] \ --shrink-factors 2x1 \ --smoothing-sigmas 1x0vox \ --transform SyN[0.1,2,0] \ --metric CC[EPI.nii,MP2RAGE.nii,1,2] \ --convergence [500x100,1e-6,10] \ --shrink-factors 2x1 \ --smoothing-sigmas 1x0vox #antsApplyTransforms -d 3 -i MP2RAGE.nii -o warped_MP2RAGE.nii -r MP2RAGE.nii-t registered_1Warp.nii.gz -t registered_0GenericAffine.mat antsApplyTransforms -d 3 -i MP2RAGE.nii -o warped_MP2RAGE.nii -r MP2RAGE.nii -t registered_1Warp.nii.gz -t registered_0GenericAffine.mat 3dcalc -a warped_MP2RAGE.nii -datum short -expr 'a' -prefix warped_MP2RAGE.nii -overwrite
- compiled from command: AntsRegistrationSyN.sh -d 3 -f EPI.nii -m MP2RAGE.nii -o registered_ -t s -n 4
- on Linux the command starts with minor “a” not capital “A”.
(for rigid+affine+syn) the 3/4 after -d means the dimension, use either 3 or 4 dimensional registration -n menas the number of cores
More infomration about ANTS details here: https://github.com/ANTsX/ANTs/wiki/Anatomy-of-an-antsRegistration-call
3.) Run Freesurfer on warped_MP2RAGE.nii
module load freesurfer #export FREESURFER_HOME=/Applications/freesurfer source $FREESURFER_HOME/SetUpFreeSurfer.sh pfad=$(pwd) export SUBJECTS_DIR=$pfad recon-all -s subject_name -hires -i warped_MP2RAGE.nii -all -parallel -openmp 64.) SUMA
- Get AFNI stuff
- in subject folder: “cd subject_name” Run @SUMA_Make_Spec_FS -sid subject_name –NIFTI
- Scale up EPI:
- in NeuroDebian: FFT EPI.nii 0 and FFT MP2RAGE.nii 0
cd subject_name @SUMA_Make_Spec_FS -sid subject_name -NIFTI cd SUMA cp ../../EPI.nii ./ cp ../../warped_MP2RAGE.nii ./ echo "************* upscaling EPI.nii ******************************" module load afni delta_x=$(3dinfo -di EPI.nii) delta_y=$(3dinfo -dj EPI.nii) delta_z=$(3dinfo -dk EPI.nii) sdelta_x=$(echo "(($delta_x / 4))"|bc -l) sdelta_y=$(echo "(($delta_x / 4))"|bc -l) sdelta_z=$(echo "(($delta_z / 4))"|bc -l) echo "$sdelta_x" echo "$sdelta_y" echo "$sdelta_z" 3dresample -dxyz $sdelta_x $sdelta_y $sdelta_z -rmode Li -overwrite -prefix scaled_EPI.nii -input EPI.nii #get obliquity matrix 3dWarp -card2oblique EPI.nii -verb warped_MP2RAGE.nii -overwrite > orinentfile.txt echo "dense mesh starting" #get dense mesh MapIcosahedron -spec subject_name_lh.spec -ld 564 -prefix std_lh.ld564. -overwrite MapIcosahedron -spec subject_name_rh.spec -ld 564 -prefix std_rh.ld564. -overwrite echo "************************ get surfaces in oblique orientation left" ConvertSurface -xmat_1D orinentfile.txt -i std_lh.ld564.lh.pial.gii -o std_lh.ld564.lh.pial.obl.gii -overwrite ConvertSurface -xmat_1D orinentfile.txt -i std_lh.ld564.lh.smoothwm.gii -o std_lh.ld564.lh.smoothwm.obl.gii -overwrite #get spec for the new file quickspec -tn gii std_lh.ld564.lh.pial.obl.gii mv quick.spec std_lh.ld564.lh.pial.obl.spec quickspec -tn gii std_lh.ld564.lh.smoothwm.obl.gii mv quick.spec std_lh.ld564.lh.smoothwm.obl.spec inspec -LRmerge std_lh.ld564.lh.smoothwm.obl.spec std_lh.ld564.lh.pial.obl.spec -detail 2 -prefix std_BOTH.ld564.lh.orient.spec -overwrite echo " **************************" echo " get binary mask of surface left" echo " **************************" 3dSurf2Vol -spec std_lh.ld564.lh.pial.obl.spec -surf_A std_lh.ld564.lh.pial.obl.gii -map_func mask -gridset scaled_EPI.nii -prefix lh.pial.epi_vol.nii -sv T1.nii -overwrite 3dSurf2Vol -spec std_lh.ld564.lh.smoothwm.obl.spec -surf_A std_lh.ld564.lh.smoothwm.obl.gii -map_func mask -gridset scaled_EPI.nii -prefix lh.WM.epi_vol.nii -sv T1.nii -overwrite 3dSurf2Vol -spec std_BOTH.ld564.lh.orient.spec -surf_A std_lh.ld564.lh.smoothwm.obl.gii -surf_B std_lh.ld564.lh.pial.obl.gii -sv T1.nii -gridset scaled_EPI.nii -map_func mask -f_steps 40 -f_index points -f_p1_fr 0.07 -f_pn_fr -0.05 -prefix ribbonmask_564_lh.nii -overwrite # is fill should be bigger #3dSurf2Vol -spec std_BOTH.ld564.lh.orient.spec -surf_A std_lh.ld564.lh.smoothwm.obl.gii -surf_B std_lh.ld564.lh.pial.obl.gii -sv T1.nii -gridset scaled_EPI.nii -map_func mask -f_steps 40 -f_index points -f_p1_fr -0.05 -f_pn_fr 0.05 -prefix ribbonmask_564_lh.nii -overwrite echo " **************************" echo " *******DONE WITH LEFT HEMISHPERE" echo " **************************" echo "************************ get surfaces in oblique orientation left" ConvertSurface -xmat_1D orinentfile.txt -i std_rh.ld564.rh.pial.gii -o std_rh.ld564.rh.pial.obl.gii -overwrite ConvertSurface -xmat_1D orinentfile.txt -i std_rh.ld564.rh.smoothwm.gii -o std_rh.ld564.rh.smoothwm.obl.gii -overwrite #get spec for the new file quickspec -tn gii std_rh.ld564.rh.pial.obl.gii mv quick.spec std_rh.ld564.rh.pial.obl.spec quickspec -tn gii std_rh.ld564.rh.smoothwm.obl.gii mv quick.spec std_rh.ld564.rh.smoothwm.obl.spec inspec -LRmerge std_rh.ld564.rh.smoothwm.obl.spec std_rh.ld564.rh.pial.obl.spec -detail 2 -prefix std_BOTH.ld564.rh.orient.spec -overwrite echo " **************************" echo " get binary mask of surface right" echo " **************************" 3dSurf2Vol -spec std_rh.ld564.rh.pial.obl.spec -surf_A std_rh.ld564.rh.pial.obl.gii -map_func mask -gridset scaled_EPI.nii -prefix rh.pial.epi_vol.nii -sv T1.nii -overwrite 3dSurf2Vol -spec std_rh.ld564.rh.smoothwm.obl.spec -surf_A std_rh.ld564.rh.smoothwm.obl.gii -map_func mask -gridset scaled_EPI.nii -prefix rh.WM.epi_vol.nii -sv T1.nii -overwrite 3dSurf2Vol -spec std_BOTH.ld564.rh.orient.spec -surf_A std_rh.ld564.rh.smoothwm.obl.gii -surf_B std_rh.ld564.rh.pial.obl.gii -sv T1.nii -gridset scaled_EPI.nii -map_func mask -f_steps 40 -f_index points -f_p1_fr 0.07 -f_pn_fr -0.05 -prefix ribbonmask_564_rh.nii -overwrite #3dLocalstat -nbhd 'SPHERE(0.2)' -prefix filled_ribbonmask_564 ribbonmask_564+orig 3dcalc -a ribbonmask_564_rh.nii -b ribbonmask_564_lh.nii -expr 'a + b ' -prefix fill.nii -overwrite 3dcalc -a lh.pial.epi_vol.nii -b rh.pial.epi_vol.nii -expr 'a + b ' -prefix pial_vol.nii -overwrite 3dcalc -a lh.WM.epi_vol.nii -b rh.WM.epi_vol.nii -expr 'a + b ' -prefix WM_vol.nii -overwrite 3dLocalstat -nbhd 'SPHERE(0.3)' -stat mean -overwrite -prefix filled_fill.nii fill.nii 3dcalc -a filled_fill.nii -b fill.nii -expr 'step(step(a-0.5)+b)' -overwrite -prefix filled_fill.nii 3dcalc -a filled_fill.nii -b pial_vol.nii -c WM_vol.nii -expr 'step(a-b-c)' -overwrite -prefix GM_robbon4_manual_corr.nii 3dcalc -a filled_fill.nii -b pial_vol.nii -c WM_vol.nii -expr 'a + b + 2*c ' -prefix rim_auto.nii -overwrite cp filled_fill.nii ../../ cp pial_vol.nii ../../ cp WM_vol.nii ../../ cp rim_auto.nii ../../ cp scaled_EPI.nii ../../ cp GM_robbon4_manual_corr.nii ../../5. manual correction of GM_ribbon.nii
fslview:
- uncheck protection
- use Pen to draw GM AREAS that are miss classified
- Use errasor to remouve GM (almost never happens)
- opposite sides of a sulcus should not touch each other
Alternatively in fsleyes:
- alt+E gets you in edit mode, where you can manually correct the GM ribbon

6.) calculating layers from GM-ribbon in neuroDebian
- Getting layers from GM-ribbon in NeuroDebian:
-
./PREP4MANrim pial_vol.nii WM_vol.nii GM_robbon4_manual_corr.nii
- outpout is rim.nii
-
- Generate Layers from rim.nii
-
- script_2run_bock_layers.sh, which basely does:
LAYER_VOL_LEAK rim.nii GROW_LAYERS rim.nii3dcalc -a leak_vol_lay_rim.nii -b equi_dist_layers.nii -expr 'a-b' -overwrite -prefix difference.nii SMinMASK difference.nii rim.nii 30 3dcalc -a smoothed_difference.nii -b leak_vol_lay_rim.nii -expr 'b-2*a' -overwrite -prefix corrected_leak_1.nii SMinMASK corrected_leak_1.nii rim.nii 12 GLOSSY_LAYERS smoothed_corrected_leak_1.nii
-
Input:  Output:
Output:
Hi, I am running the tutorial to extract layers from the GM-ribbon with the ultimate goal of extracting layer-specific values of functional connectivity (from VASO acquisition), T1 relaxation time from MP2RAGE, and QSM. I successfully ran the majority of the steps but cannot find in the LAYNII downloads the commands /PREP4MANrim and script_2run_bock_layers.sh. How can I download the files? It is written elsewhere that it requires Renzo’s Neurodebian virtual box but the link does not work. Are there alternative commands to perform the same? Thanks a lot for your help.
Cheers,
Giovanni
LikeLike
Dear Giovanni,
Thanks for your interest in this pipeline. Unfortunately, this pipeline is quite old by now. The PREP4MANrim (https://github.com/layerfMRI/repository/tree/master/ODIN_related/PREP4MANrim) was part of the ODIN-dependent version of LayNii and is no longer supported.
I believe that the more current features in LayNii might get you what you want:
E.g. see LN2_BORDERIZE https://github.com/layerfMRI/LAYNII/blob/master/src/LN2_BORDERIZE.cpp
see LN2_RIMIFY https://github.com/layerfMRI/LAYNII/blob/master/src/LN2_RIMIFY.cpp
or see the corresponding scripts here: https://github.com/ofgulban/LAYNII_extras/tree/master/RIMIFY and here: https://github.com/ofgulban/LAYNII_extras/tree/master/get_rim_from_FARUKsegm
If you need help with a spefific problem and/or dataset, feel free to post an issue here on github with a link to the data and someone will help you: https://github.com/layerfMRI/LAYNII/issues/new
Best regards,
Renzo
LikeLike