Authors: Ömer Faruk Gülban, Renzo Huber | Date: April 2024
Chapter published here: https://www.sciencedirect.com/science/article/abs/pii/B9780128204801001881
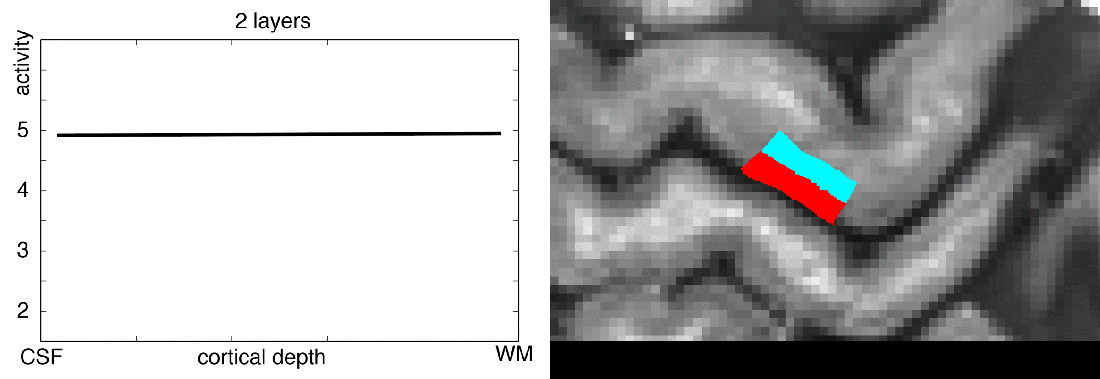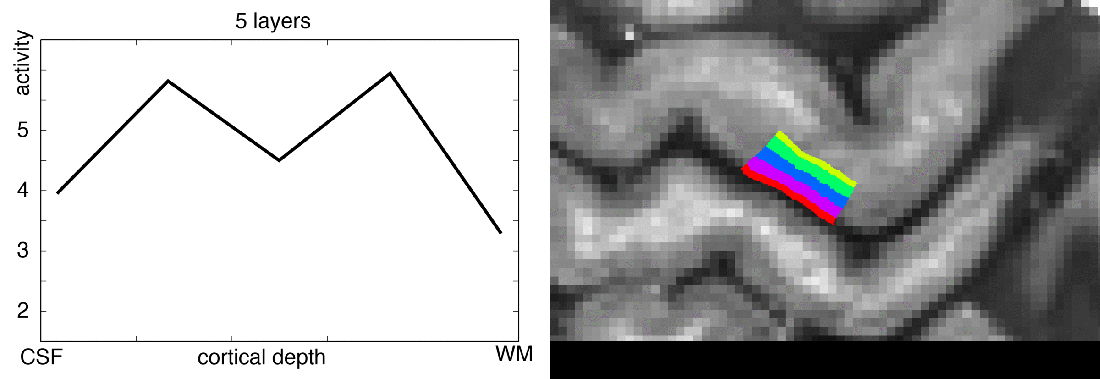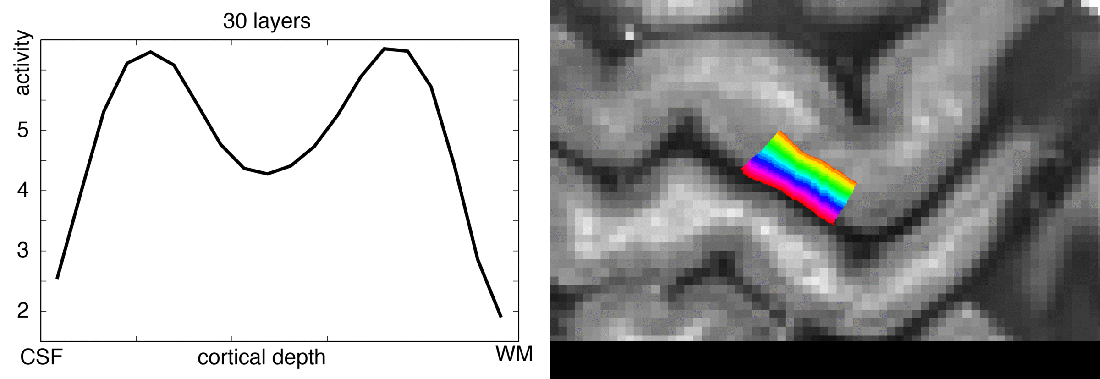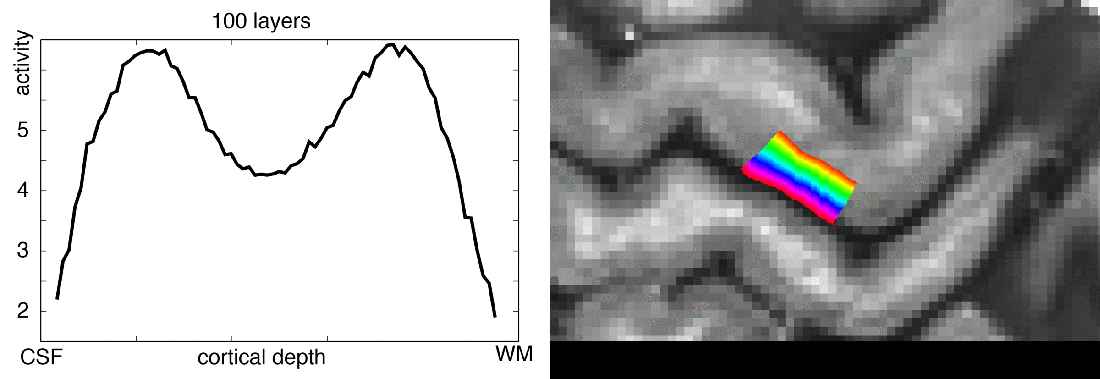
Functional magnetic resonance imaging (fMRI) today is a common method to study the human brain. The popularity of fMRI can be explained by its versatility and accessibility to study the structure-function relationship in living humans, which has historically been challenging or impossible. One area of constant progress and excitement since the early days in fMRI is the advances in hardware, MR sequences, and software which allowed researchers to image the fine details of the human brain non-invasively. fMRI is currently capable of reaching sub-millimeter resolution, effectively transforming the MRI scanner from a macroscope into a mesoscope. While 0.8 mm isotropic voxel resolution (~0.5 μl) has become a part of the daily routine of a high resolution fMRI researcher at 7 Tesla scanners, as opposed to e.g. 3 mm isotropic voxels in conventional fMRI studies conducted at 3 T (Bollmann et al. 2020), recent developments have breached 0.37 mm isotropic resolution (~0.05 μl) (Feinberg et al. 2023). This 10 fold reduction in voxel volume promises an exciting future to observe the fine mesoscopic details such as cortical layers, columns, and vessels during changes in human brain function.
Continue reading “Computing geometric layers and columns on continuously improving human (f)MRI data”