In this blog post, I want to describe the application and working principle of a few spatial smoothing algorithms that are implemented in LAYNII.
- Confined smoothing along similar anatomical structures with
LN_GRADSMOOTH - Smoothing along the layers with
LN_LAYERSMOOTH - Smoothing within columns with
LN_LAYERSMOOTH - Smoothing across specific spatial dimensions only with
LN_DIRECT_SMOOTH - Smoothing long the time domain with
LN_TEMPSMOOTH
Introduction
I have mixed feelings about spatial smoothing.
On one hand, it is paradoxical to go through all the trouble of acquiring high-resolutions data, when the spatial information will be later lost during spatial smoothing anyway. My former co-adviser Bob Turner is known (and feared) for repeatedly pointing out the disadvantages of smoothing. Thus, my reluctance to spatial smoothing has become second nature to me even before I had really understood what smoothing actually was.
On the other hand, smoothing can really improve the sensitivity of tSNR-starved high-resolution fMRI-data. Thus, I cannot always resist the temptation to apply some sort of spatial smoothing. In this blog post, I want to discuss potential smoothing algorithms that do not necessarily reduce the spatial specificity too much.
1.) Spatial smoothing within similar anatomical compartments only
In many studies, one is interested in the activity of various layers in a given patch of gray matter. And any signal leakage and smoothing signal from the pial vasculature (in CSF space) might compromise the results in gray matter. In this scenario, it is beneficial to apply spatial smoothing within brain compartments only (e.g. within tissue types or layers). This kind of spatially informed spatial smoothing can be done in the LAYNII program LN_GRADSMOOTH. LN_GRADSMOOTH, has two inputs, 1) the 3D or 4D image that should be smoothed and 2) a reference image with some sort of anatomical contrast. The anatomical reference image does not need be pre-segmented in anyway and can be bases on any available contrast (phase, T2*, tSNR, T1 etc.). Based on this reference image, smoothing is only applied across voxels that share a similar anatomical signal in the reference image.
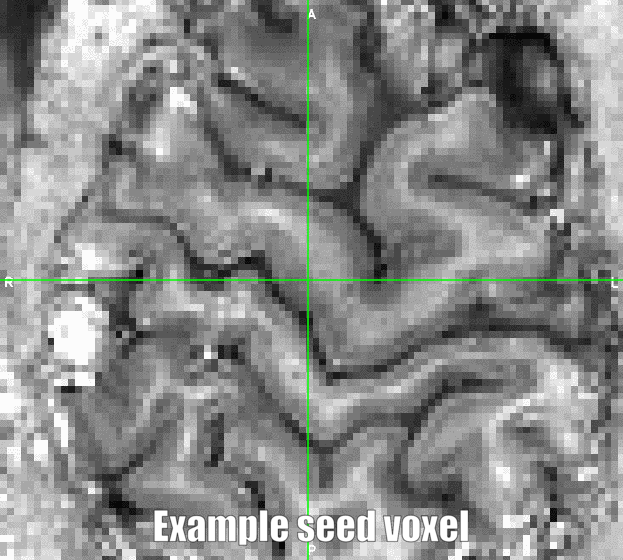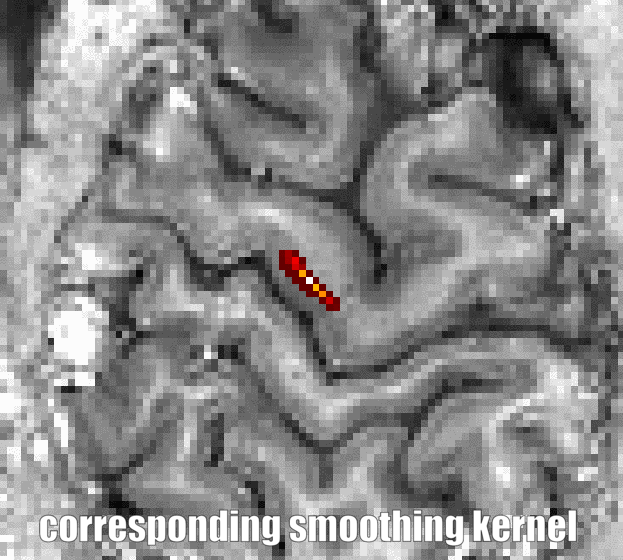
The program LN_GRADSMOOTH can be downloaded here and executed as follows:
LN_GRADSMOOTH -gradfile T1.nii -input timeseries.nii -FWHM 1 -within -selectivity 0.08, whereas the -selectivity parameter determines how similar the voxels need to be in order to be included in the same smoothing kernel. E.g. a value of 0.5 smoothes within tissue classes (GM, CSF) only, whereas a value of 0.08 smoothes within layers only.

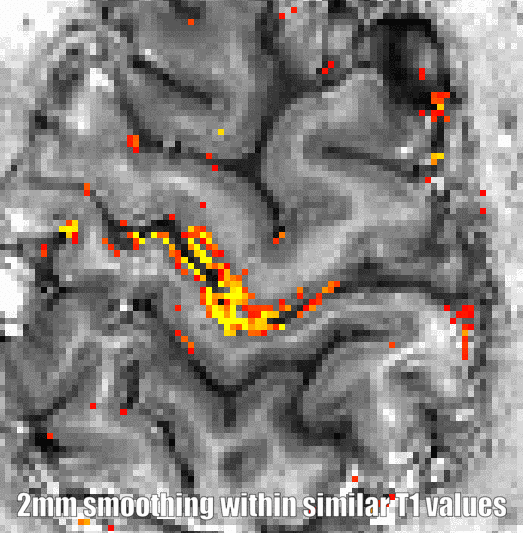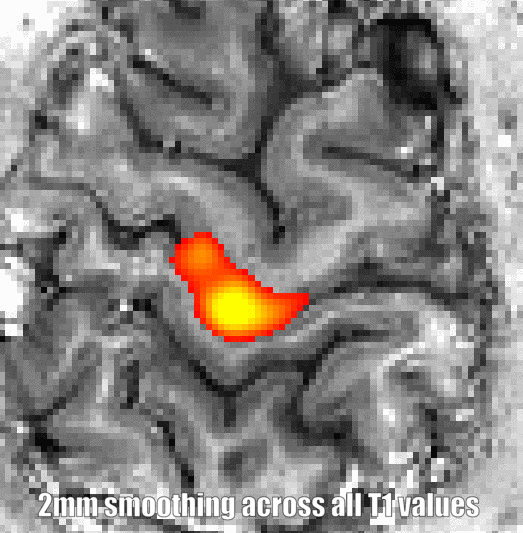
There are two fundamental advantaged of this kind of smoothing compared to the other smoothing algorithms below: Firstly, is does not require any ground truth anatomical clarification input. As such it is not necessary to have a high-quality segmentation, estimated of cortical depths, topologically consistent geometry etc. Secondly, it can be done in the native EPI resolution and does not require upscaling to a finer grid, as it is necessary for proper layer estimations. Thus, it can easily be applied on time-series without making file sizes unnecessary big.
One potential disadvantage is that it relies on the assumption that similar functional units have similar anatomical signal intensities. While this assumption is somewhat valid across tissue types such as GM/WM/CSF, and across cortical depths, it is violated for columnar fMRI and across GM across opposite cortical banks.
The signal intensity informed smoothing described here might be somewhat similar to the FSL smoothing algorithm SUSAN, with the important distinction that SUSAN can only exploit the spatial structure in the data that are supposed to be smoothed, not allowing the usage of an external reference.
2.) Smoothing within layers
More accurate layer smoothing can be done with LN_LAYER_SMOOTH, which has been described in this previous blog post. The underlying algorithm is basically an iterative smoothing within a given (layer) mask only.

This way of smoothing can be very useful for to better visualize layer signatures in activity maps.
The program can be downloaded here and executed as:
LN_LAYER_SMOOTH -layer_file layers.nii -input activity_map.nii -FWHM 1
When the smoothing kernel is too big (E.g. larger than 1 mm), it can happen that this procedure results in leakage from across a sulcus. E.g. that the signal in the upper layer of the sensory cortex is smoothed together with the signal in the upper layer of the motor cortex, just because they are right across from each other in the central sulcus.

To account for such cases, there is the -NoKissing option. When this option is selected, smoothing is only applied along connected layers within one side of GM. Unconnected areas that are are close to each other in Euclidean space, are ignored.

-NoKissing option avoids this.The advantage of this smoothing method is that it is anatomically much more accurate that the smoothing algorithm in 1.). However, it requires an input of predefined layer estimates, which can be cumbersome to generate. Furthermore, this smoothing is best executed on a finer spatial grid than the native EPI resolution. Thus, the resulting file size can be quite big. Especially for 4D time series data.
3.) Smoothing within columns
In studies, where the columnar (not the laminar) organization principle of the cortex is of interest, it can be beneficial to apply spatial smoothing perpendicular to the cortical layers only. This columnar smoothing can also be done with the program LN_LAYER_SMOOTH.

The program can be downloaded here and is executed as:
LN_LAYER_SMOOTH -layer_file layers.nii -input activity_map.nii -FWHM 1
The columnar masks can be generated as described here.
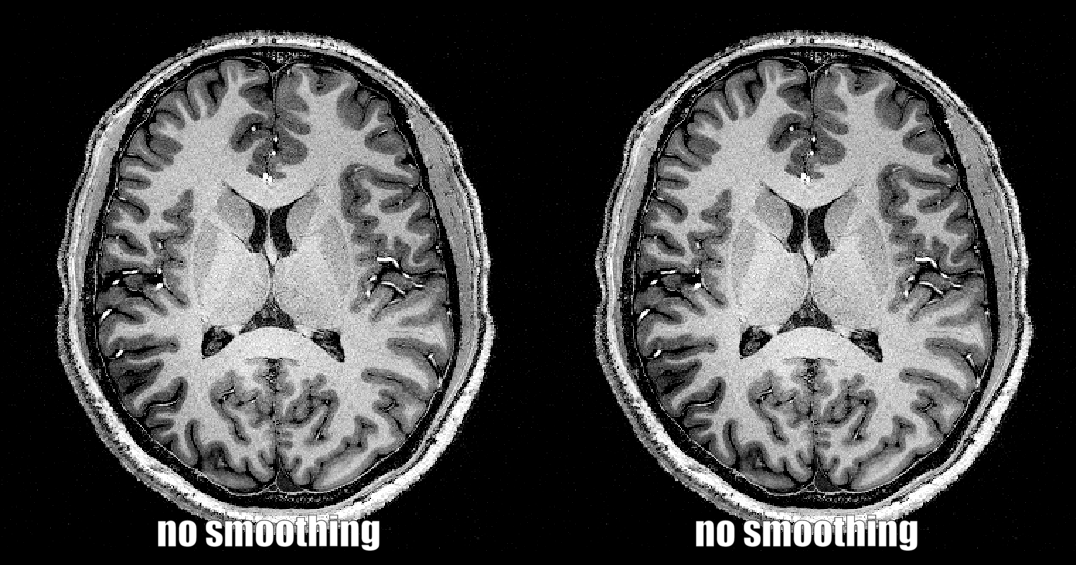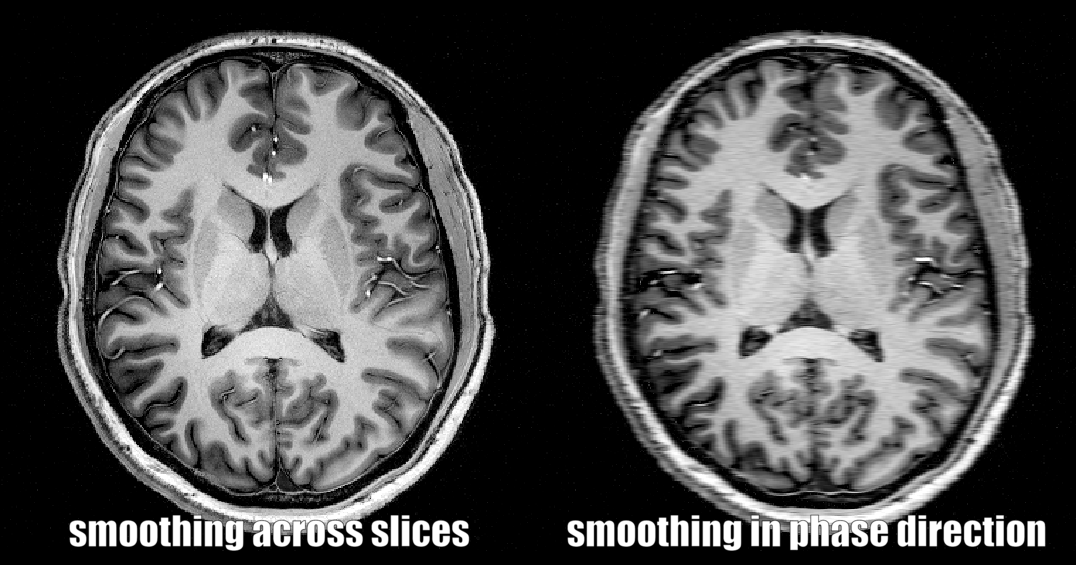
4.) Smoothing across specific spatial dimensions only
In some instances, it can make sense to restrict the spatial smoothing certain spatial dimensions only.
- E.g. smoothing only in the phase encoding direction to simulate T2* blurring,
- E.g. smoothing only in the slice dimension to see if how the data would look like with thicker slices.
- E.g. smoothing along the slab dimension to visualice large vessel pathways across slices (similar to mean intensity projection).
- E.g. smoothing across layers or columns in rectangular IMAGIRO data.
This kind of smoothing is implemented in LN_DIRECT_SMOOTH, which can be dowloaded here and is executed as:
LN_DIRECT_SMOOTH -input activity_map.nii -FWHM 1 -direction 1
5.) Smoothing along time dimension
For fMRI time series, it can often be helpful to apply smoothing in the temporal dimension. I found it for example helpful with resting-state data, where one cannot increase the contrast to noise ratio by averaging across trials.
This kind of temporal smoothing can also be referred to as low pass filtering (removing high frequencies).
In the LAYNII program LN_TEMPSMOOTH there are two ways of temporal filtering implemented
- Temporal smoothing with a gaußian smoothing kernel of a given FWHM.
LN_TEMPSMOOTH -timeseries file.nii -gaus 1.0 - Temporal smoothing with a box car smoothing kernel of a given width. This is basically a sliding window averaging.
LN_TEMPSMOOTH -timeseries file.nii -box 1.0


Scripts of doing similar things other software suites are here for FSL.
Comment on LAYNII versions
This blog post is written and tested for LAYNII version v1.5.6. Earlier versions might required the flag –sulctouch, instead of –NoKissing.