Did you acquire your data with GE-BOLD EPI and now worry about draining vein contaminations? There are several models out there for post-processing that should help you to tease out the tiny microvascular GE-BOLD signal that you care about and help you to remove the dominating macro-vascular venous signal. However, note that some of these vein-removal models work better than others. None of the models is perfect! But some of them are useful. The most relevant approaches are implemented in the LAYNII software suit on a voxel-wise level.
In this blog post, I want to describe these de-veining models and how to use them to get rid of unwanted macrovascular venous signals in LAYNII.
Background
In layer-fMRI with GE-BOLD, the majority of the signal (80%-90% at 7T) is coming from layer unspecific draining veins. This is particularly apparent in superficial layers. Close to the pial surface, only a small part of the signal is actually interesting to us.
Lacking better acquisition methods than GE-BOLD, many clever researchers came up with a number of layer-dependent BOLD signal models that describe this signal bias towards the superficial layers. The aim of these models is usually not only to characterize the macrovascular bias and to understand its origin, but they usually go beyond that. Often these models are ultimately aiming for the removal of the unwanted vein signal, while retaining the microvasculature signal of interest.
There are three classes of models that received some attention in the field. These models are implemented in LAYNII and are discussed below.
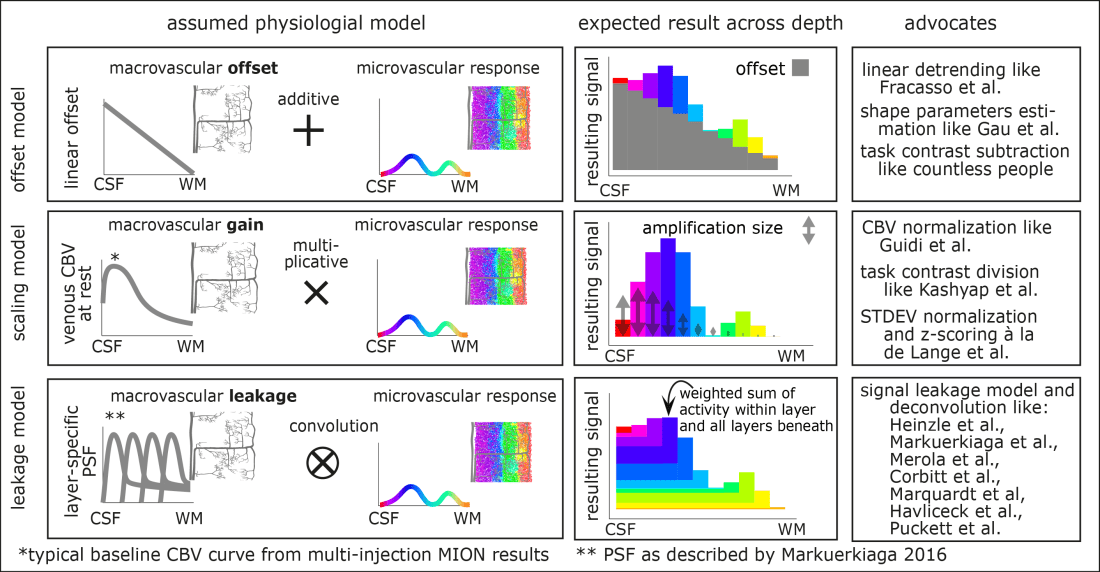
All of these models aim to distinguish the unwanted macrovascular signal from the desired microvascular response. And all of these models have in common that the macrovascular bias contributes a much stronger signal (80%-90%) compared to the microvascular response (10%-20%). Furthermore, all of these models predict the superficial bias in layer-dependent GE-BOLD data. However, the individual models differ in the way how the macrovascular signal is assumed to come about. In the different models, the origin of the macrovascular bias is assumed to be different.
The linear-offset model is based on the assumption that the layer-specific microvascular signal is added on top of a task-independent macrovascular signal. Thus, it is assumed that a simple task-contrast subtraction or a simple linear-trend-removal would get rid of the unwanted macrovascular component of the BOLD signal.
The scaling model is based on the assumption that the layer-specific bias in GE-BOLD is driven by layer-dependent variations of vein density. As such, it is assumed that the superficial signal is larger than the signal in deeper layers, simply because the amount of venous blood volume is higher in superficial layers compared to deeper layers. Or in other words, the macrovasculature acts like a layer-specific signal amplification (gain). Thus, it is assumed that a simple multiplicative (or divisory) normalization can correct for the macrovascular signal bias. This layer-dependent signal normalization has been proposed as part of multiple layer-fMRI analysis strategies, including:
- (a) scaling the layer-dependent signal with estimates of layer-dependent venous CBV,
- (b) normalizing the signal difference between different task responses by the mean signal response of all involved task responses,
- (c) refraining to infer neuroscience conclusions solely based on task response differences, instead focusing on task response ratios.
The leakage model is based on the assumption that the BOLD signal in each layer constitutes a signal mixture of neural activity from multiple layers. Namely, it is assumed that a voxel in a given layer contains some neural signal from the layer of interest plus an unwanted integrated activity signal of all the layers beneath. E.g. it is assumed that the BOLD signal in a voxel of a superficial layer contains both, the desired superficial signal and unwanted signal from the deeper layers. The BOLD signal model in each layer is then usually parameterized as a weighted sum of the layer itself and this signal from deeper layers. The corresponding layer-dependent weight factors of the weighted sum can be estimated with more-or-less plausible physiological assumptions about the underlying layer-dependent blood vessel architecture. To ultimately use this model to correct for unwanted leaked signal, a spatial signal deconvolution approach is applied. (However, note that some implementations of such deconvolution models refrain from the term “deconvolution” in favor of the terms ‘matrix inversion” or “spatial signal subtraction”). Very often, the leakage model is combined with the scaling model.
In most layer-dependent application papers to date, these models are applied on an area-wide scale. I.e. usually fMRI signals are pooled from the layers of large cortical patches and their effect is solely investigated in the form of one-dimensional layer profiles. In LAYNII, however, the various deveining algorithms are implemented to work on a voxel-by-voxel level. This allows the researcher to appreciate the signal distribution along in the topographical space across layers and columns. Furthermore, the resulting signal maps provide an intuitive understanding of the noise amplification that comes along with the various de-veining algorithms of the respective models.
Example data set to exemplify different de-veining models in LAYNII
The algorithms of vein removal and their application in LAYNII is exemplified by means of the test data set that comes along with the LAYNII software on Github. The example data can be downloaded from the test_data folder on Github and include:
lo_layers.nii, lo_T1EPI.nii, lo_columns.nii, lo_ALF.nii, lo_BOLD_act.nii, lo_VASO_act.nii
- The layer file lo_layers.nii provides an integer value for each voxel in the cortical depth. You can generate your own layer file for your own data set in LAYNII with LN_GROW_LAYERS as explained on this tutorial here.
- The column file lo_columns.nii provides integer values for each cortical distance along the cortical ribbon in voxel-space. It can be generated in LAYNII with LN_3DCOLUMNS as explained on this tutorial here.
- The voxel-wise estimate of venous baseline blood volume (CBVv) lo_ALF.nii can be generated by means of the resting-state fluctuation amplitude (RSFA) or from the task residuals by means of the VASA contrast (a.k.a ALF). For the purpose of layer-dependent vein removal, it is not necessary that this estimate is given in quantitative units nor is it necessary that it is if free of bias fields contaminations across the FOV. It only needs to have consistent units on a column-by-column scale. It is extremely simple to generate such an estimate of (CBVv). Namely, one just needs to estimate the temporal variance of slow signal fluctuations. This can be done with LN_TEMPSMOOTH in LAYNII, or in FSL with this script (more information on the physiological basis and underlying assumptions is given below).
- The BOLD file lo_BOLD_act.nii refers to a tapping induced signal change of 30sec on-off paradigm. Note that this file has the largest signal change in the superficial layers. The LAYNII program LN2_DEVEIN aims to remove this bias.
- The T1-EPI file lo_T1EPI.nii and the VASO file lo_VASO_act.nii are included just for reference. They are not used in any algorithm.

Option 1: Linear trend removal across cortical depth
The linear trend removal in LN2_DEVEIN is applied when the -linear flag is used. The algorithm simply uses the depth information provided in the lo_layers.nii file to remove the superficial bias.

This approach of linear trend removal has been advocated and applied on one-dimensional layer-profile level by Fracasso et al. and Gau et al. However, this approach is also implicitly used when researchers try to remove the superficial signal bias by subtracting BOLD responses of different task conditions from each other.
Example application of linear layer-detrending in LAYNII with a –linear flag
LN2_DEVEIN -linear -layer_file lo_layers.nii -input lo_BOLD_act.nii
Option 2: Normalization based on known distributions of layer-dependent baseline venous blood volume
The CBV scaling removal in LN2_DEVEIN is applied when the -CBV flag is used. The algorithm simply normalizes the depth-dependent BOLD signal in each layer and column with the vascular reactivity measure given in the lo_ALF.nii file. Since the venous baseline CBV is larger in superficial layers, this normalization removes the surface bias in the BOLD response map.

This approach is often applied in layer-fMRI studies, when researchers investigate the relative difference of task conditions as opposed to absolute differences. Furthermore, this approach is also explicitly used when researchers normalize multiple task responses by each other on a layer-by-layer level. The physiological basis of BOLD signal scaling has been advocated by Kazan et al., Kashyap et al., and developed further for single task condition normalization in Guidi et al.
The layer dependent normalization by the signal variance or so called “z-scoring” is based on similar assumptions. With the small distinction that the z-scoring is not specific to venous CBV scaling, but also might add a bias of thermal and CSF noise into the layer scaling too.
Example application of layer-dependent CBVv scaling in LAYNII with the -CBV flag
LN2_DEVEIN -CBV -layer_file lo_layers.nii -column_file lo_columns.nii -input lo_BOLD_act.nii -ALF lo_ALF.nii
Option 3: Layer-dependent vascular deconvolution based on signal leakage models
Descriptions of the BOLD signal by means of spatial signal leakage across layers has been proposed and discussed by a large number of research labs, including models from Heinzle et al., Markuerkiaga et al., Merola et al., Puckett et al., Corbitt et al., Marquardt et al., and Havliceck et al. All of these signal leakage models estimate the signal in superficial layers as a sum of the microvascular response within the given layer plus a weighted sum of the signal from all other layers below. The major difference between different leakage models comes from the procedure of how the respective summation weights are estimated. Usually the summation weights are derived based on more-or-less appropriate assumptions of the layer-dependent blood vessel architecture and/or experimental data. The weights are usually dependent on resting CBVv and CBF assumptions across cortical depth.

Model assumptions in the voxel-wise deconvolution de-veining approach
The vascular deconvolution model is the most sophisticated among the three models discussed here. It is also the model that best resembles the signal origin as far as it is understood. However, while it is the physiologically most plausible model, its implementation in LAYNII relies upon higher order assumptions still:
- The depth-dependent deconvolution weights are relatively stable across time and are not changing faster than the time scale of TR. I tested this assumption by comparing the voxel-wise de-veined BOLD signal with simultaneously acquired VASO (as ground truth) in movie watching tasks and resting-state data. In those data, the functional contrast cannot be collapsed across TRs within long stimulus blocks. Instead, every single TR resembles its own stimulus condition. As far as I looked, I could not see any violation of this assumption. However, I believe that at hypothetical sub-second TRs, the time delay between layers due to finite blood flow velocities between layers will violate this assumption.
- There is no tangential venous signal leakage across columns. I.e. it is assumed that each column is drained from deeper layers towards superficial layers only. Tangential blood drainage is not considered. Thus, the application of the LAYNII deconvolution results might be hard to interpret at the spatial scale of the tangential venous branch length (<0.6mm). This assumption will not be violated if the columnar units in the column file are kept to be thicker than approx. 0.6mm-0.8mm. In case you have thinner columnar estimates, note that the thickness of the estimated columnar units can be adjusted in LAYNII with the -Ncolumns and the -vinc flag, respectively.
- In the LAYNII implementation, there is no assumption on layer-dependent venous blood volume necessary. While most implementations of layer-fMRI leakage models need to assume the underlying venous CBV at rest, this is not necessary in the implementation in LAYNII. This can be quite advantageous because, as discussed previously (e.g. by Markuerkiaga and Havliceck) the local baseline CBVv has a large effect on the deconvolution weights. And thus, a small error in the assumed CBVv will result in a large error of the de-veined BOLD results. Since CBVv varies a lot along the cortical sheet (e.g. between neighboring columns), this is a big limitation for voxel-wise applications of column-specific deconvolution approaches. To avoid this potential source of error, the ALF dataset is given as an input to LAYNII. This allows LAYNII to perform the de-veining on a column-by-column level. Ultimately, the only assumption remains that the ALF is a good estimate of CBVv. I have validated this extensively across voxels and across cortical depths by means of hypercapnia in multiple publications here and here.
- The layer-dependent baseline CBF is within a reasonable order of magnitude. As discussed previously (e.g. by Markuerkiaga) different assumptions about the baseline CBF do not really affect the deconvolution weights a lot, nor do they strongly affect the de-veined BOLD results. Thus, a large error in the assumed baseline CBF results in a small error of the deconvolved results. As discussed by Markuerkiaga et al., even these small effects can be accounted for by a scaling parameter that adjusts the peak-to-tail ratio of the layer-dependent point spread function. In LN2_DEVEIN, this can be adjusted by means of an optional -lambda parameter. If you need to assume that your CBF is exceptionally low, use -lambda 0.3. If you need to assume that your CBF is exceptionally high, use -lambda 0.2.

When one assumes very large or very low CBF values, the resulting de-veined BOLD signal maps are not affected a lot.
Example application of layer-dependent vein-deconvolution in LAYNII
LN2_DEVEIN -layer_file lo_layers.nii -column_file lo_columns.nii -input lo_BOLD_act.nii -ALF lo_ALF.nii
All program executions used in this blog post are tested for LAYNII version v.1.5.6 and will probably work for later versions too.
Acknowledgements
I would like to thank Kamil Uludag for countless discussions about layer-fMRI models and their application: it was a great July 2019. Thanks also to Kamil for completing the list of deconvolution models cited here. I would like to thank Faruk Gulban for his contributions to LAYNII (2.0) that made it much more straightforward to write and document LN2_DEVEIN.